John Gurdon
Distinguished group leaderResearch summary
Nuclear reprogramming by oocytes and eggs
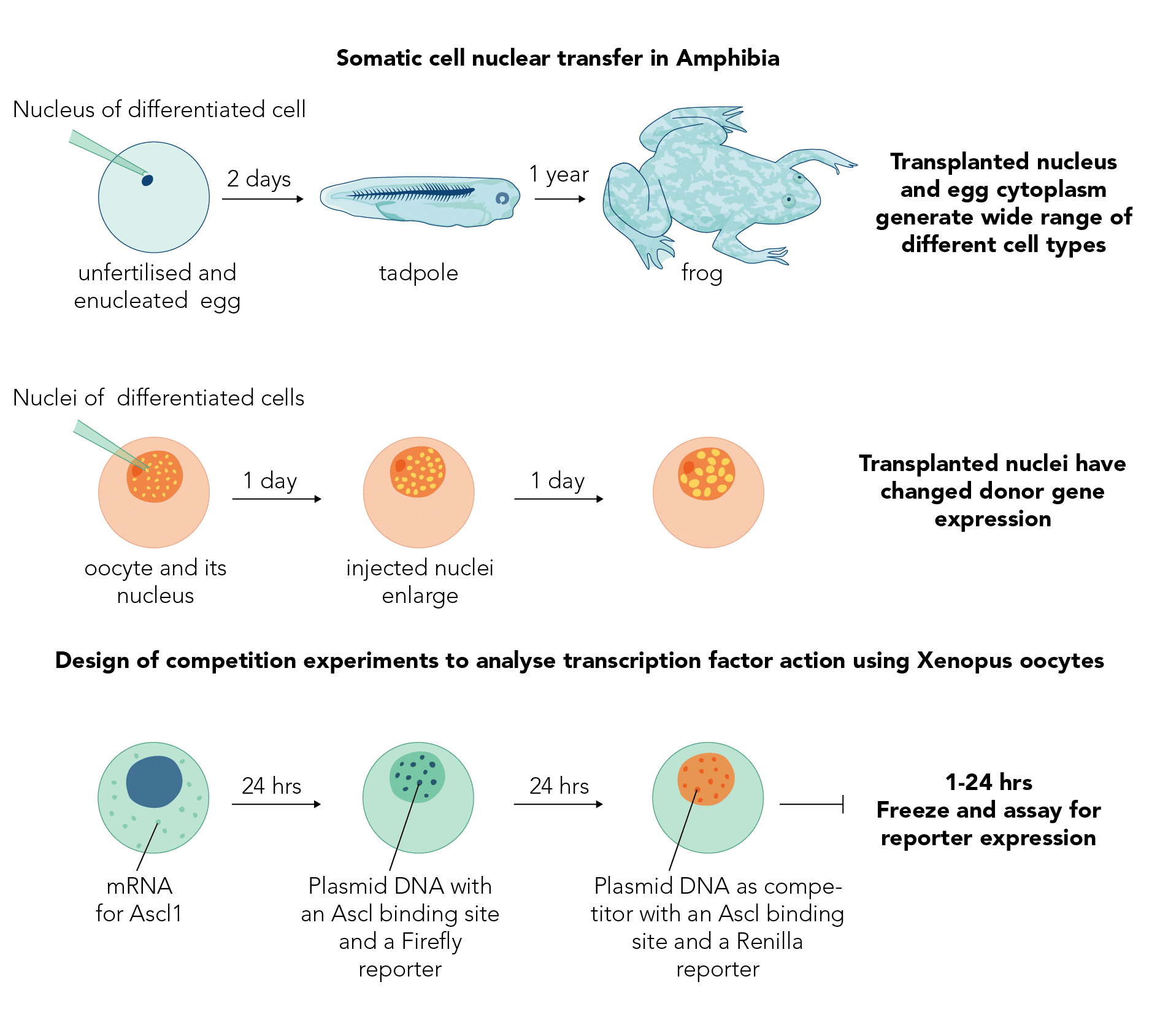
Can we make cell reprogramming more efficient? Our group focuses on somatic cell nuclear transfer to amphibian eggs and oocytes from two complementary points of view. One aims to identify the molecules and mechanisms by which the cytoplasm of an egg or oocyte can reprogramme the nucleus of a differentiated somatic cell to behave like that of an embryo. From this state, many different kinds of cells for replacement can be generated.

Selected publications
-
Gurdon JB et al. (2020) Long-term association of a transcription factor with its chromatin binding site can stabilize gene expression and cell fate commitment. Proc Natl Acad Sci USA 117: 15075–15084. DOI: 10.1073/pnas.2000467117.
-
Aztekin C et al. (2019) Identification of a regeneration-organizing cell in the Xenopus tail. Science 364: 653–658. DOI: 10.1126/science.aav9996.
-
Hörmanseder E et al. (2017) H3K4 methylation-dependent memory of somatic cell identify inhibits reprogramming and development of nuclear transfer embryos. Cell Stem Cell 21: 135–143.e6. DOI: 10.1016/j.stem.2017.03.003.
-
Gurdon J (2017) Nuclear transplantation, the conservation of the genome, and prospects for cell replacement. FEBS J 284(2): 211-217. DOI: 10.1111/febs.13988.
Biography
Professor Sir John Gurdon Kt DPhil DSc FRS
Distinguished Group Leader, Nobel Laureate in Physiology or Medicine 2012, Member of the University Department of Zoology
Educated at Eton College, where he did Classics, having been advised that he was unsuited for science, and Christ Church, Oxford (Zoology). PhD with Michael Fischberg, on nuclear transplantation in Xenopus. Obtained the first clone of genetically identical adult animals. Demonstrated genetic totipotency of somatic cell nuclei by obtaining sexually mature frogs from the nuclei of intestinal epithelium.
Did postdoctoral work at Cal-Tech, on bacteriophage genetics. Moved to MRC Molecular Biology Laboratory in Cambridge (Chairman Max Perutz), subsequently becoming Head of Cell Biology Division. In 1983, accepted John Humphrey Plummer Professorship of Cell Biology in University of Cambridge, in Zoology Department. Initiated, with Prof R Laskey, Cancer Research Campaign unit of Molecular Embryology in Zoology Department Cambridge.
In 1990 moved to new Wellcome CRC Institute of Cancer and Developmental Biology in Cambridge, and served as Chairman 1990-2001. From 2001, the Institute was renamed The Gurdon Institute. Master of Magdalene College, Cambridge 1995 to 2002, and Governor (= Trustee) of the Wellcome Trust 1995 to 2000. Chairman of the Company of Biologists from 2001 to 2011.
Dr Gurdon has received multiple awards and recognitions internationally, too numerous to list them all here (we provide a selection, and more can be found in ‘Who’s Who’).
Notable achievements and honours
-
2017Golden Plate Award, US Academy of Achievement
-
2016Gold Medal from Indian PM Narendra Modi
-
2012Nobel Prize for Physiology or Medicine
-
2009Albert Lasker Award for Basic Medical Research
-
2003Copley Medal, Royal Society
-
1995Knight Bachelor
-
1989Wolf Prize in Medicine
-
1985Royal Medal, Royal Society
-
1984Prix Charles Leopold Mayer, Academie des Sciences, France