Iva Tchasovnikarova
Group leaderResearch summary
Genetic interrogation of epigenetic pathways
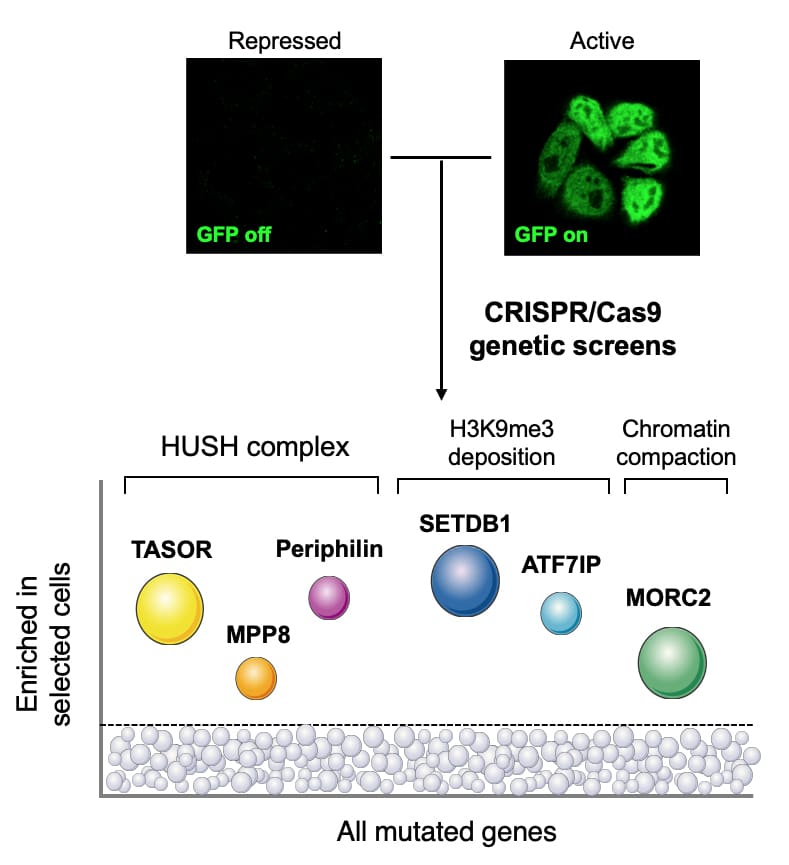
We study epigenetic pathways and the mechanisms through which these processes are corrupted by disease-associated mutations in chromatin regulators. We aim to (1) understand the mechanisms utilised by chromatin modifiers to exert their function in healthy human cells, and (2) examine how these mechanisms are altered in human disorders in molecular detail.

Selected publications
-
Danac JM et al. (2024) Competition between two HUSH complexes orchestrates the immune response to retroelement invasion. Molecular Cell 84, 1–12. DOI: 10.1016/j.molcel.2024.06.020.
-
Tchasovnikarova IA et al. (2021) TRACE generates fluorescent human reporter cell lines to characterize epigenetic pathways. Molecular Cell 82: 479–491.e7. DOI: 10.1016/j.molcel.2021.11.035.
-
Douse CH et al. (2020) TASOR is a pseudo-PARP that directs HUSH complex assembly and epigenetic transposon control. Nature Communications, 11: 4940. DOI: 10.1038/s41467-020-18761-6.
-
Guillen Sacoto MJ et al. (2020) De Novo Variants in the ATPase Module of MORC2 Cause a Neurodevelopmental Disorder with Growth Retardation and Variable Craniofacial Dysmorphism. Am J Hum Gen 107: 352-363. DOI: 10.1016/j.ajhg.2020.06.013.
-
Timms RT et al. (2019) Differential Viral Accessibility (DIVA) identifies alterations in chromatin architecture through large-scale mapping of lentiviral integration sites. Nat Protoc 14: 153–170. DOI: 10.1038/s41596-018-0087-5.
-
Tchasovnikarova IA et al. (2017) Hyperactivation of HUSH complex function by Charcot-Marie-Tooth disease mutation in MORC2. Nat Genet 49: 1035–1044. DOI:10.1038/ng.3878.
-
Tchasovnikarova IA et al. (2015) Epigenetic silencing by the HUSH complex mediates position-effect variegation in human cells. Science 348: 1481–1485. DOI: 10.1126/science.aaa7227.
Biography
Iva Tchasovnikarova PhD
Group Leader, Member of the University Department of Biochemistry
Originally from Bulgaria, Iva moved to the UK to study Biotechnology at the University of Edinburgh. She then joined the laboratory of Prof. Paul Lehner at the Cambridge Institute for Medical Research at the University of Cambridge, as part of the Wellcome Trust Infection and Immunity 4-year PhD programme. There, she performed a series of haploid gene-trap mutagenesis and CRISPR/Cas9-mediated forward genetic screens which identified a novel epigenetic repressor complex, HUSH, as the key mediator of position-effect variegation in human cells.
In 2016, Iva joined the laboratory of Dr Robert Kingston at Massachusetts General Hospital/Harvard Medical School as a fellow of the Damon Runyon Cancer Research Foundation. There, she leveraged her experience with genetic technologies to develop a genome-wide method to study chromatin accessibility (DIVA), as well as an unbiased method that allows for the identification of fluorescent, cell-based reporters of chromatin states (TRACE). Dr Tchasovnikarova joined the Gurdon Institute as a Group Leader in 2020.
Notable achievements and honours
-
2020Damon Runyon-Dale F. Frey Award for Breakthrough Scientists
-
2016Damon Runyon Cancer Research Foundation Postdoctoral Fellowship
Research group
-
Joshua Danac
PhD Student
-
Rachael Matthews
PhD Student
-
Chuyan Qin
PhD Student
-
Dr Akhila Gungi
Research Associate