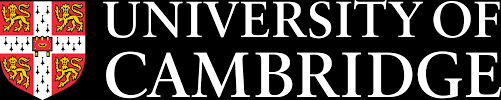Azim Surani
Director of germline and epigenetics researchResearch summary
The human germline
We study primordial germ cells (PGCs), precursors to eggs and sperm, in the early embryo. We have established principles of early human development with a focus on human PGC (hPGC) specification. A unique epigenetic resetting follows in the germline after hPGC specification. Our work shows that SOX17 is the key regulator of human, but not mouse, germ cell fate. By developing in vitro models, and with authentic hPGCs from human embryos, we have also established how pluripotent stem cells gain competence for germ cell and somatic fates in human.

Selected publications
-
Kobayashi T et al. (2021) Tracing the emergence of primordial germ cells from bilaminar disc rabbit embryos and pluripotent stem cells. Cell Reports 37(2):109812. DOI: 10.1016/j.celrep.2021.109812.
-
Sybirna A et al. (2020) A critical role of PRDM14 in human primordial germ cell fate revealed by inducible degrons. Nat Commun 11: 1282. DOI: 10.1038/s41467-020-15042-0.
-
Hackett JA et al. (2018) Tracing the transitions from pluripotency to germ cell fate with CRISPR screening. Nat Commun 9(1): 4292. DOI: 10.1038/s41467-018-06230-0.
-
Floros VI et al. (2018) Segregation of mitochondrial DNA heteroplasmy through a developmental genetic bottleneck in human embryos. Nat Cell Biol 20: 144– 151. DOI: 10.1038/s41556-017-0017-8.
-
Kobayashi T et al. (2017) Principles of early human development and germ cell program from conserved model systems. Nature 546: 416–420. DOI: 10.1038/nature22812.
-
Murakami K et al. (2016) NANOG alone induces germ cells in primed epiblast in vitro by activation of enhancers. Nature 529: 403–407. DOI: 10.1038/nature16480.
-
Tang WWC et al. (2015) A Unique Gene Regulatory Network Resets the Human Germline Epigenome for Development. Cell 161(6): 1453–1467. DOI: 10.1016/j.cell.2015.04.053.
-
Irie N et al. (2015) SOX17 Is a Critical Specifier of Human Primordial Germ Cell Fate. Cell 160: 253–268. DOI: 10.1016/j.cell.2014.12.013.
Biography
Prof Azim Surani PhD CBE FRS FMedSci
Director of Germline and Epigenetics Research, Wellcome Senior Investigator, Member of the University Department of Physiology, Development and Neuroscience
The discovery of genomic imprinting in 1984 by Surani was important for advances in mammalian development and the field of epigenetics. Mammalian genomes show epigenetic symmetry in totipotent zygotes because the chromosomes retain a memory of their parental origin with heritable DNA methylation tags, which regulate monoallelic expression of parental alleles of ‘imprinted’ genes. These genes have a role in mammalian development, growth, and behaviour; aberrant imprints underlie some human diseases.
Notably, Surani also elucidated the genetic basis for mammalian primordial germ cell specification in mice and humans, which initiates the unique mammalian germline epigenetic program, including erasure and reestablishment of parental imprints. His continuing research is on early human development, the origin of germ cells during gastrulation, and epigenetic inheritance.
Surani was born in Kenya and received a PhD at Cambridge University under Sir Robert Edwards (Nobel laureate, 2010) in 1975, and was elected as Marshall-Walton Professor in 1992, and then as Director of Germline and Epigenetics Research in 2013 at the Gurdon Institute.
He is a Fellow of the Royal Society and the Academy of Medical Sciences. His awards include the William Bate Hardy Prize, a Royal Medal for mammalian development, Rosenstiel Award for epigenetic regulation of gene expression in mammalian embryos, ISSCR McEwen Award for Innovation, and the Canada Gairdner International Award for genomic imprinting and epigenetics.
Notable achievements and honours
-
2022Mendel Medal, Genetics Society
-
2018Canada Gairdner International Award
-
2017Linacre Lecture (St John’s College, Cambridge)
-
2014McEwen Award for Innovation, International Society for Stem Cell Research
-
2014Jawaharlal Nehru Fellowship, Government of India
-
2010Galton Lecture, The Galton Institute
-
2010Royal Medal, Royal Society
-
2010Mendel Lecture (Brno, Czech Republic)
-
2007Commander of the British Empire for services to biology
-
2007Lewis S Rosenstiel Award for Distinguished Work in Basic Medical Science
-
2004Sir Dorabji Tata Distinguished Professor, NCBS Institute (Bangalore) and Distinguished Fellow, Nehru Center
-
2001Fellow of the Academy of Medical Sciences
-
1999Wellcome-Burroughs Research Pioneer Award, National Institute of Child Health and Human Development (USA)
-
1994Doctor of Science honoris causa, University of Uppsala (Sweden)
-
1994Member of Academia Europaea
-
1993Member of European Molecular Biology Organisation
-
1992Associate Fellow of the Third World Academy of Sciences
-
1986 & 1992BBSRC Individual Merit Award
-
1991William Bate Hardy Prize, Cambridge Philosophical Society
-
1990Menten Visiting Professor of Pathology, University of Pittsburgh (USA)
-
1990Fellow of the Royal Society
Research group
-
Lynn Froggett
Lab Administrator
-
Dr Theresa Gross-Thebing
Visiting Scholar
-
Dr Wolfram Gruhn
Research Associate
-
Dr Mei Gu
Research Assistant/ Lab Manager
-
Dr Geraldine Jowett
Research Associate
-
Dr Jitesh Neupane
Research Associate
-
Dr So Shimamoto
Visiting Scholar
-
Dr Baojiang Wu
Visiting Scholar
-
Carmela Cruz
Visiting Student
-
Alexander Karpovitsch
Visiting Student