Deciphering Notch’s targets and mechanism
April 25, 2022
Read more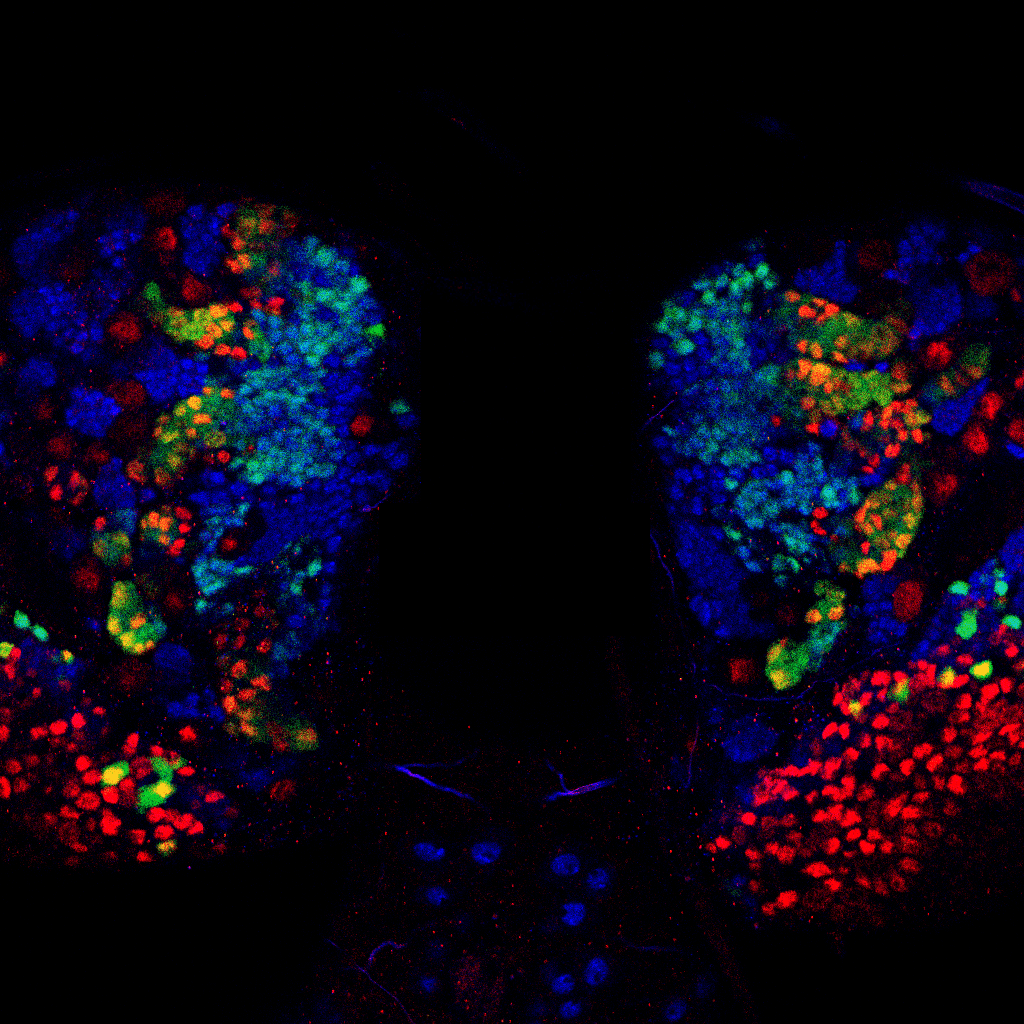
A recent study from the Brand lab led by Jocelyn Tang, Anna Hakes and Robert Krautz has employed a new technique – NanoDam – to identify previously unknown temporal factors involved in generating neuronal diversity in the Drosophila brain and visual system. The technique enables rapid cell type specific, genome-wide profiling of protein-DNA interactions with high temporal and spatial resolution.
Tang LY et al. (2022) NanoDam identifies Homeobrain (ARX) and Scarecrow (NKX2.1) as conserved temporal factors in the Drosophila central brain and visual system. Developmental Cell 57(9): 1193-1207.E7. DOI: 10.1016/j.devcel.2022.04.008.
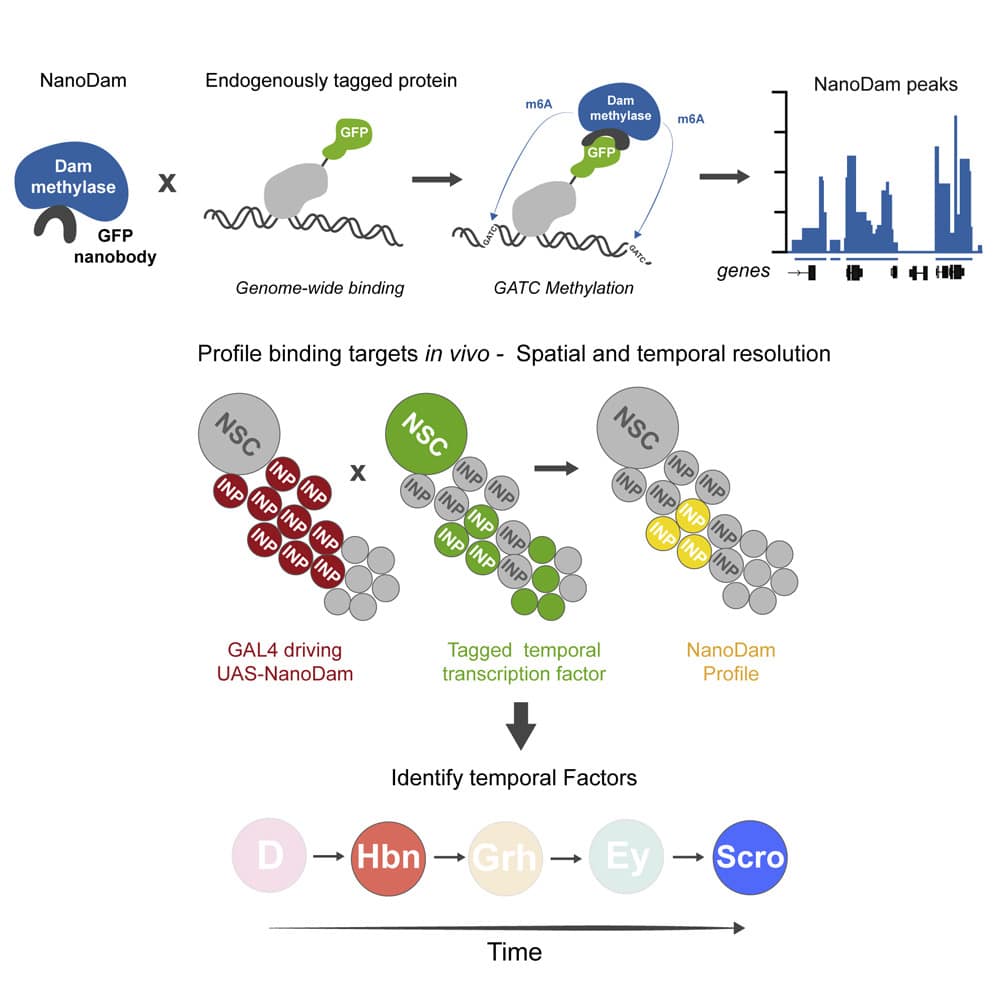
Temporal patterning of neural progenitors is an evolutionarily conserved strategy for generating neuronal diversity. Type II neural stem cells in the Drosophila central brain produce transit-amplifying intermediate neural progenitors (INPs) that exhibit temporal patterning. However, the known temporal factors cannot account for the neuronal diversity in the adult brain.
To search for missing factors, we developed NanoDam, which enables rapid genome-wide profiling of endogenously tagged proteins in vivo with a single genetic cross. Mapping the targets of known temporal transcription factors with NanoDam revealed that Homeobrain and Scarecrow (ARX and NKX2.1 orthologs) are also temporal factors. We show that Homeobrain and Scarecrow define middle-aged and late INP temporal windows and play a role in cellular longevity. Strikingly, Homeobrain and Scarecrow have conserved functions as temporal factors in the developing visual system.
NanoDam enables rapid cell-type-specific genome-wide profiling with temporal resolution and is easily adapted for use in higher organisms.