Deciphering Notch’s targets and mechanism
April 25, 2022
Read more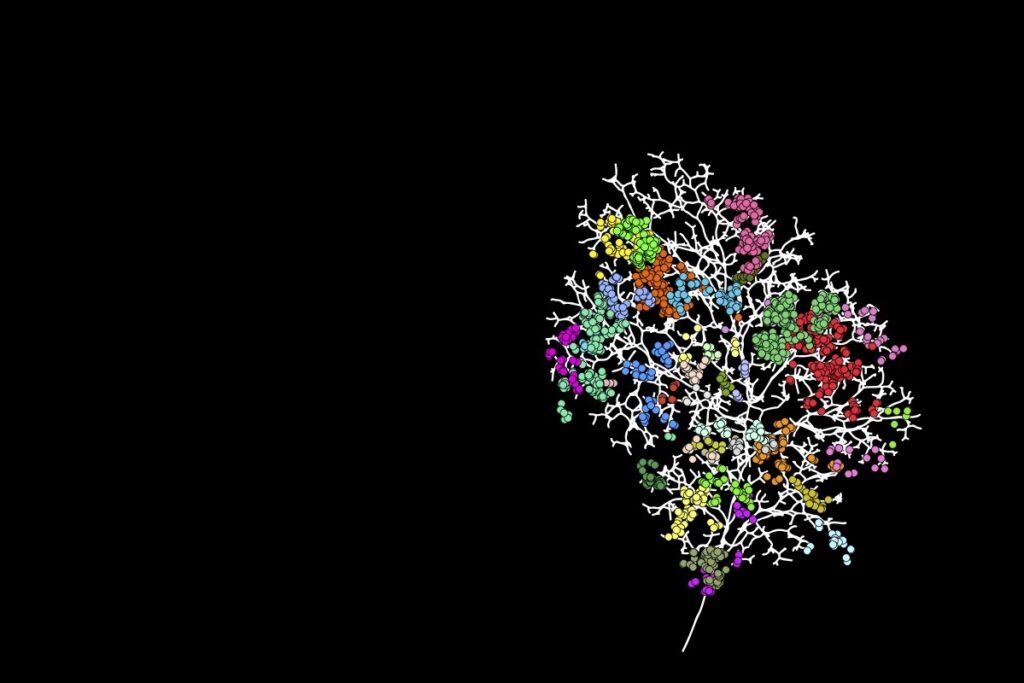
A study from the Simons lab led by Lemonia Chatzeli and Ignacio Bordeu showed how cell fate specification correlates with position biased and epithelial patterning of the mouse salivary gland epithelium. The lab developed a system where mathematical modelling, cell lineage tracing, and single-cell RNA-seq analysis can be used to understand branching morphogenesis in combination with lineage decision making.
Chatzeli L, Bordeu I, et al. (2023) A cellular hierarchy of Notch and Kras signaling controls cell fate specification in the developing mouse salivary gland. Dev Cell. 2023 Jan 23;58(2):94-109.e6. DOI: 10.1016/j.devcel.2022.12.009.
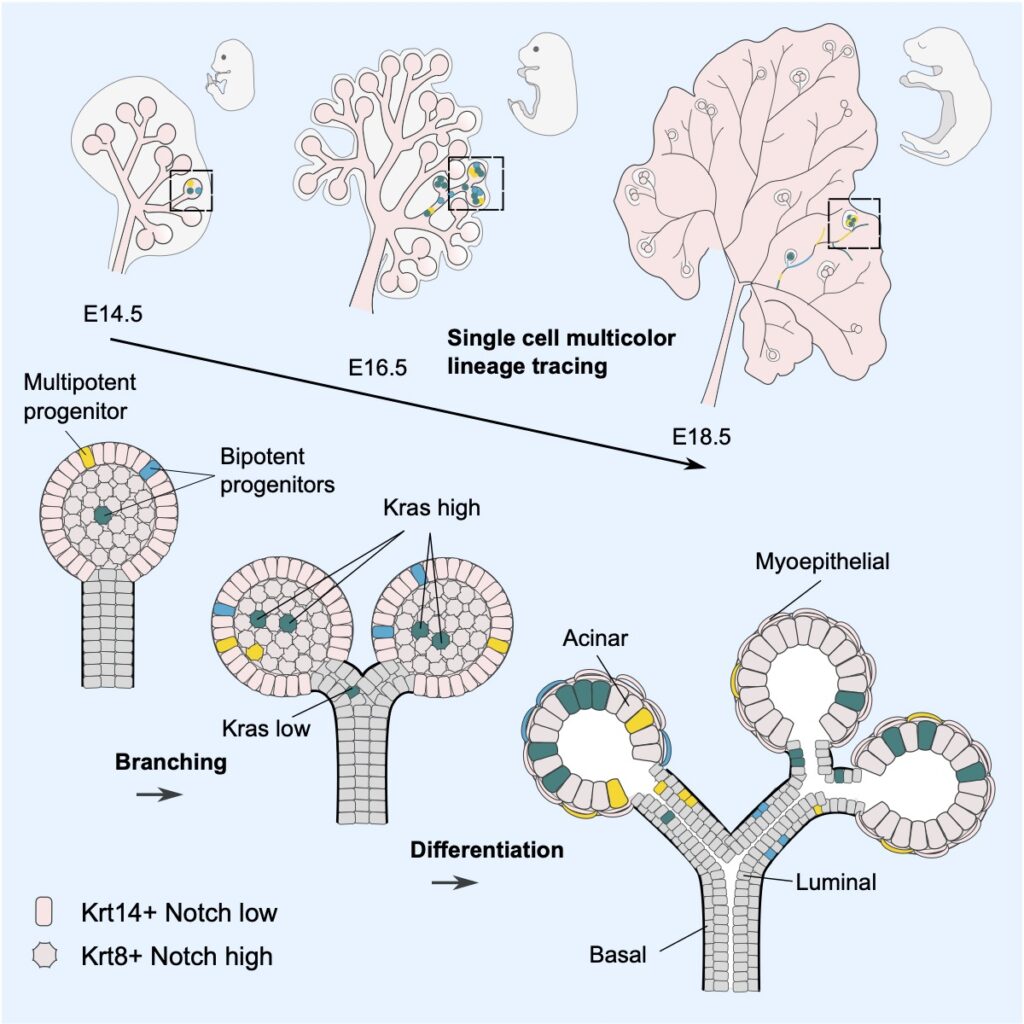
The development of the mouse salivary gland involves a tip-driven process of branching morphogenesis that takes place in concert with differentiation into acinar, myoepithelial, and ductal (basal and luminal) sub-lineages. By combining clonal lineage tracing with a three-dimensional (3D) reconstruction of the branched epithelial network and single-cell RNA-seq analysis, we show that in tips, a heterogeneous population of renewing progenitors transition from a Krt14+ multipotent state to unipotent states via two transcriptionally distinct bipotent states, one restricted to the Krt14+ basal and myoepithelial lineage and the other to the Krt8+ acinar and luminal lineage. Using genetic perturbations, we show how the differential expression of Notch signaling correlates with spatial segregation, exits from multipotency, and promotes the Krt8+ lineage, whereas Kras activation promotes proacinar fate. These findings provide a mechanistic basis for how positional cues within growing tips regulate the process of lineage segregation and ductal patterning.