How polarised epithelial cells grow “up” without breaking the barrier
September 28, 2022
Read more
This atlas provides a foundational dataset that researchers can use for generating and testing hypotheses for how cell differentiation occurs in lung development.
The team used the atlas to make predictions about progenitor cell states, signalling interactions and lineage-defining transcription factors, and demonstrated how these can be efficiently tested using a genetically-tractable human fetal lung organoid model. In addition, they discovered a new cell type that does not exist in mice and is related to a sub-type of human small cell lung cancer.
He P et al. (2022) A human fetal lung cell atlas uncovers proximal-distal gradients of differentiation and key regulators of epithelial fates. Cell 185 (25), 4841-4860.e25. DOI: 10.1016/j.cell.2022.11.005.
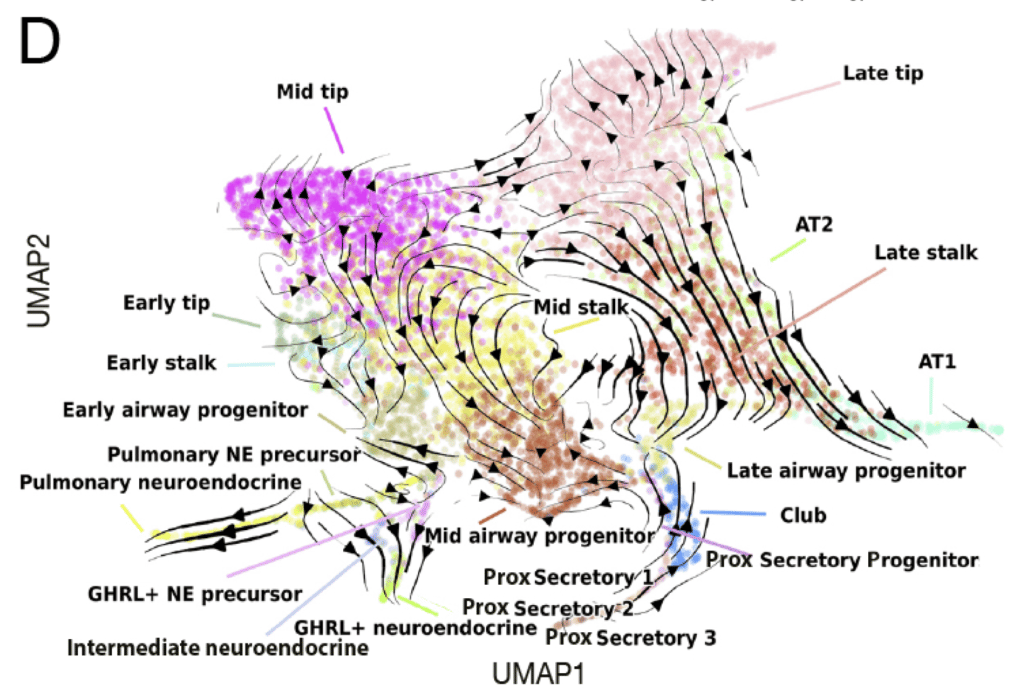
Visualisation of the predicted epithelial cell lineage trajectory
We present a multiomic cell atlas of human lung development that combines single cell RNA and ATAC sequencing, high throughput spatial transcriptomics and single cell imaging. Coupling single cell methods with spatial analysis has allowed a comprehensive cellular survey of the epithelial, mesenchymal, endothelial and erythrocyte/leukocyte compartments from 5-22 post conception weeks. We identify new cell states in all compartments. These include developmental-specific secretory progenitors and a new subtype of neuroendocrine cell related to human small cell lung cancer. Our datasets are available through our web interface. To illustrate its general utility, we use our cell atlas to generate predictions about cell-cell signalling and transcription factor hierarchies which we test using organoid models.